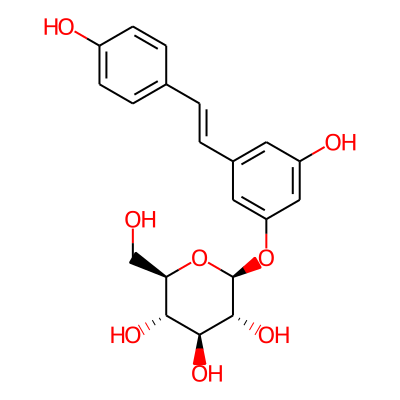
Summary
SMILES: OC[C@H]1O[C@@H](Oc2cc(/C=C/c3ccc(cc3)O)cc(c2)O)[C@@H]([C@H]([C@@H]1O)O)OInChI: InChI=1S/C20H22O8/c21-10-16-17(24)18(25)19(26)20(28-16)27-15-8-12(7-14(23)9-15)2-1-11-3-5-13(22)6-4-11/h1-9,16-26H,10H2/b2-1+/t16-,17-,18+,19-,20-/m1/s1InChIKey: HSTZMXCBWJGKHG-CUYWLFDKSA-N
DeepSMILES: OC[C@H]O[C@@H]Occc/C=C/cccccc6))O)))))))ccc6)O)))))))[C@@H][C@H][C@@H]6O))O))O
Scaffold Graph/Node/Bond level: C(=Cc1cccc(OC2CCCCO2)c1)c1ccccc1
Scaffold Graph/Node level: C1CCC(CCC2CCCC(OC3CCCCO3)C2)CC1
Scaffold Graph level: C1CCC(CCC2CCCC(CC3CCCCC3)C2)CC1
Functional groups: CO; c/C=C/c; cO; cO[C@@H](C)OC
Chemical classification
ClassyFire Kingdom: Organic compounds
ClassyFire Superclass: Phenylpropanoids and polyketidesClassyFire Class: Stilbenes
ClassyFire Subclass: Stilbene glycosides
NP Classifier Biosynthetic pathway: Shikimates and Phenylpropanoids
NP Classifier Superclass: Stilbenoids
NP Classifier Class: Monomeric stilbenes
Synonymous chemical names:piceid, piceid (resveratrol-3-o-beta-d-glucoside)
External chemical identifiers:CID:5281718; ChEMBL:CHEMBL142652; ChEBI:8198; ZINC:ZINC000004098633; FDASRS:XM261C37CQ; SureChEMBL:SCHEMBL41411; MolPort-003-937-179
Chemical structure download