Summary
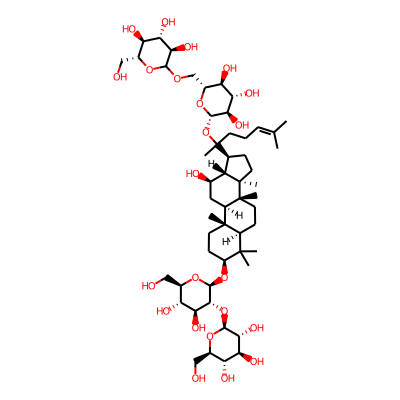
SMILES: OC[C@H]1O[C@@H](O[C@H]2CC[C@]3([C@H](C2(C)C)CC[C@@]2([C@@H]3C[C@@H](O)[C@H]3[C@@]2(C)CC[C@@H]3C(O[C@@H]2O[C@H](COC3O[C@H](CO)[C@H]([C@@H]([C@H]3O)O)O)[C@H]([C@@H]([C@H]2O)O)O)(CCC=C(C)C)C)C)C)[C@@H]([C@H]([C@@H]1O)O)O[C@@H]1O[C@H](CO)[C@H]([C@@H]([C@H]1O)O)OInChI: InChI=1S/C54H92O23/c1-23(2)10-9-14-54(8,77-48-44(69)40(65)37(62)29(74-48)22-70-46-42(67)38(63)34(59)26(19-55)71-46)24-11-16-53(7)33(24)25(58)18-31-51(5)15-13-32(50(3,4)30(51)12-17-52(31,53)6)75-49-45(41(66)36(61)28(21-57)73-49)76-47-43(68)39(64)35(60)27(20-56)72-47/h10,24-49,55-69H,9,11-22H2,1-8H3/t24-,25+,26+,27+,28+,29+,30-,31+,32-,33-,34+,35+,36+,37+,38-,39-,40-,41-,42+,43+,44+,45+,46?,47-,48-,49-,51-,52+,53+,54?/m0/s1InChIKey: GZYPWOGIYAIIPV-NGBMAODDSA-N
DeepSMILES: OC[C@H]O[C@@H]O[C@H]CC[C@][C@H]C6C)C))CC[C@@][C@@H]6C[C@@H]O)[C@H][C@@]6C)CC[C@@H]5CO[C@@H]O[C@H]COCO[C@H]CO))[C@H][C@@H][C@H]6O))O))O)))))))[C@H][C@@H][C@H]6O))O))O))))))CCC=CC)C)))))C))))))))))C)))))C))))))[C@@H][C@H][C@@H]6O))O))O[C@@H]O[C@H]CO))[C@H][C@@H][C@H]6O))O))O
Scaffold Graph/Node/Bond level: C1CCC(OCC2CCCC(OCC3CCC4C3CCC3C5CCC(OC6OCCCC6OC6CCCCO6)CC5CCC43)O2)OC1
Scaffold Graph/Node level: C1CCC(OCC2CCCC(OCC3CCC4C3CCC3C5CCC(OC6OCCCC6OC6CCCCO6)CC5CCC43)O2)OC1
Scaffold Graph level: C1CCC(CCC2CCCC(CCC3CCC4C3CCC3C5CCC(CC6CCCCC6CC6CCCCC6)CC5CCC43)C2)CC1
Functional groups: CC=C(C)C; CO; COC(C)OC; CO[C@@H](C)OC
Chemical classification
ClassyFire Kingdom: Organic compounds
ClassyFire Superclass: Lipids and lipid-like moleculesClassyFire Class: Prenol lipids
ClassyFire Subclass: Terpene glycosides
NP Classifier Biosynthetic pathway: Terpenoids
NP Classifier Superclass: Triterpenoids
NP Classifier Class: Dammarane and Protostane triterpenoids
Synonymous chemical names:gypenoside liii
External chemical identifiers:CID:73148; SureChEMBL:SCHEMBL19875636
Chemical structure download