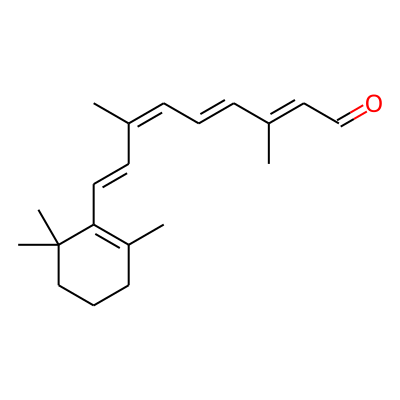

| Property name | Tool | Property value |
|---|---|---|
| Molecular weight (g/mol) | RDKit | 0 |
| Log P | RDKit | 0 |
| Topological polar surface area (Å2) | RDKit | |
| Number of hydrogen bond acceptors | RDKit | |
| Number of hydrogen bond donors | RDKit | |
| Number of carbon atoms | RDKit | |
| Number of heavy atoms | RDKit | |
| Number of heteroatoms | RDKit | |
| Number of nitrogen atoms | RDKit | |
| Number of sulfur atoms | RDKit | |
| Number of chiral carbon atoms | RDKit | |
| Stereochemical complexity | RDKit | 0 |
| Number of sp hybridized carbon atoms | RDKit | |
| Number of sp2 hybridized carbon atoms | RDKit | |
| Number of sp3 hybridized carbon atoms | RDKit | |
| Shape complexity | RDKit | |
| Number of rotatable bonds | RDKit | |
| Number of aliphatic carbocycles | RDKit | |
| Number of aliphatic heterocycles | RDKit | |
| Number of aliphatic rings | RDKit | |
| Number of aromatic carbocycles | RDKit | |
| Number of aromatic heterocycles | RDKit | |
| Number of aromatic rings | RDKit | |
| Total number of rings | RDKit | |
| Number of saturated carbocycles | RDKit | |
| Number of saturated heterocycles | RDKit | |
| Number of saturated rings | RDKit | |
| Number of Smallest Set of Smallest Rings (SSSR) | RDKit |

| Property name | Tool | Property value |
|---|---|---|
| Number of Lipinski’s rule of 5 violations | RDKit | 1 |
| Lipinski’s rule of 5 filter | RDKit | Passed |
| Number of Ghose filter violations | RDKit | 1 |
| Ghose filter | RDKit | Failed |
| Veber filter | RDKit | Good |
| Pfizer 3/75 filter | RDKit | Bad |
| GSK 4/400 filter | RDKit | Bad |
| Weighted quantitative estimate of drug-likeness (QEDw) score | RDKit | 0.3585 |

| Property name | Tool | Property value |
|---|---|---|
| Bioavailability score | SwissADME | 0.55 |
| Solubility class [ESOL] | SwissADME | Moderately soluble |
| Solubility class [Silicos-IT] | SwissADME | Soluble |
| Blood Brain Barrier permeation | SwissADME | No |
| Gastrointestinal absorption | SwissADME | Low |
| Log Kp (Skin permeation, cm/s) | SwissADME | -4.27 |
| Number of PAINS structural alerts | SwissADME | 0.0 |
| Number of Brenk structural alerts | SwissADME | 1.0 |
| CYP1A2 inhibitor | SwissADME | No |
| CYP2C19 inhibitor | SwissADME | Yes |
| CYP2C9 inhibitor | SwissADME | Yes |
| CYP2D6 inhibitor | SwissADME | No |
| CYP3A4 inhibitor | SwissADME | No |
| P-glycoprotein substrate | SwissADME | No |

| Protein identifier | HGNC symbol | Combined score from STITCH database |
|---|---|---|
| ENSP00000249750 | ALDH1A2 | 900 |
| ENSP00000257895 | RDH5 | 962 |
| ENSP00000263126 | AKR1C4 | 700 |
| ENSP00000265605 | ALDH8A1 | 835 |
| ENSP00000267502 | RDH12 | 900 |
| ENSP00000297785 | ALDH1A1 | 970 |
| ENSP00000306606 | ADH1B | 700 |
| ENSP00000334801 | DHRS4L2 | 900 |
| ENSP00000337915 | CYP3A4 | 700 |
| ENSP00000352584 | AKR1B10 | 700 |
| ENSP00000363832 | AOX1 | 900 |
| ENSP00000369927 | AKR1C3 | 700 |
| ENSP00000370254 | AKR1C1 | 700 |