
| Property name | Tool | Property value |
|---|---|---|
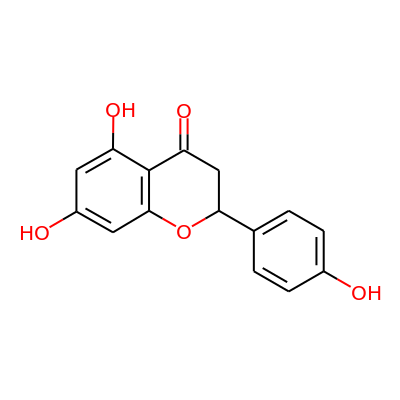
| Molecular weight (g/mol) | RDKit | 272.26 |
| Log P | RDKit | 2.51 |
| Topological polar surface area (Å2) | RDKit | 86.99 |
| Number of hydrogen bond acceptors | RDKit | 5 |
| Number of hydrogen bond donors | RDKit | 3 |
| Number of carbon atoms | RDKit | 15 |
| Number of heavy atoms | RDKit | 20 |
| Number of heteroatoms | RDKit | 5 |
| Number of nitrogen atoms | RDKit | 0 |
| Number of sulfur atoms | RDKit | 0 |
| Number of chiral carbon atoms | RDKit | 1 |
| Stereochemical complexity | RDKit | 0.07 |
| Number of sp hybridized carbon atoms | RDKit | 0 |
| Number of sp2 hybridized carbon atoms | RDKit | 13 |
| Number of sp3 hybridized carbon atoms | RDKit | 2 |
| Fraction of sp3 hybridized carbon atoms (Fsp3) | RDKit | 0.13 |
| Shape complexity | RDKit | 0.13 |
| Number of rotatable bonds | SwissADME | 1 |
| Number of aliphatic carbocycles | RDKit | 0 |
| Number of aliphatic heterocycles | RDKit | 1 |
| Number of aliphatic rings | RDKit | 1 |
| Number of aromatic carbocycles | RDKit | 2 |
| Number of aromatic heterocycles | RDKit | 0 |
| Number of aromatic rings | RDKit | 2 |
| Total number of rings | RDKit | 3 |
| Number of saturated carbocycles | RDKit | 0 |
| Number of saturated heterocycles | RDKit | 0 |
| Number of saturated rings | RDKit | 0 |
| Number of Smallest Set of Smallest Rings (SSSR) | RDKit | 3 |

| Property name | Tool | Property value |
|---|---|---|
| Number of Lipinski’s rule of 5 violations | RDKit | 0 |
| Lipinski’s rule of 5 filter | RDKit | Passed |
| Number of Ghose filter violations | RDKit | 0 |
| Ghose filter | RDKit | Passed |
| Veber filter | RDKit | Good |
| Pfizer’s 3/75 filter | RDKit | Good |
| GSK 4/400 filter | RDKit | Good |
| Number of Leadlikeness violations | SwissADME | 0 |
| Weighted quantitative estimate of drug-likeness (QEDw) score | RDKit | 0.74 |

| Property name | Tool | Property value |
|---|---|---|
| Bioavailability score | SwissADME | 0.55 |
| Solubility class [ESOL] | SwissADME | Soluble |
| Solubility class [Silicos-IT] | SwissADME | Soluble |
| Blood Brain Barrier permeation | SwissADME | No |
| Gastrointestinal absorption | SwissADME | High |
| Log Kp (Skin permeation, cm/s) | SwissADME | -6.17 |
| Number of PAINS structural alerts | SwissADME | 0 |
| Number of Brenk structural alerts | SwissADME | 0 |
| CYP1A2 inhibitor | SwissADME | Yes |
| CYP2C19 inhibitor | SwissADME | No |
| CYP2C9 inhibitor | SwissADME | No |
| CYP2D6 inhibitor | SwissADME | No |
| CYP3A4 inhibitor | SwissADME | Yes |
| P-glycoprotein substrate | SwissADME | Yes |

| Protein identifier | HGNC symbol | Combined score from STITCH database |
|---|---|---|
| ENSP00000216117 | HMOX1 | 816 |
| ENSP00000225831 | CCL2 | 816 |
| ENSP00000228434 | CD69 | 800 |
| ENSP00000229794 | MAPK14 | 800 |
| ENSP00000233242 | APOB | 822 |
| ENSP00000260010 | TLR2 | 800 |
| ENSP00000260433 | CYP19A1 | 776 |
| ENSP00000260630 | CYP1B1 | 817 |
| ENSP00000262186 | KCNH2 | 800 |
| ENSP00000262735 | PPARA | 860 |
| ENSP00000265724 | ABCB1 | 825 |
| ENSP00000308227 | HMGA1 | 800 |
| ENSP00000316329 | SCD5 | 733 |
| ENSP00000320709 | ADIPOQ | 800 |
| ENSP00000332049 | CD86 | 786 |
| ENSP00000335657 | CCK | 800 |
| ENSP00000342007 | CYP1A2 | 822 |
| ENSP00000358414 | HMGCS2 | 784 |
| ENSP00000359380 | SCD | 800 |
| ENSP00000361264 | RAPGEF1 | 800 |
| ENSP00000385806 | KCNMA1 | 786 |
| ENSP00000414303 | BDNF | 800 |
| ENSP00000454071 | LDLR | 800 |